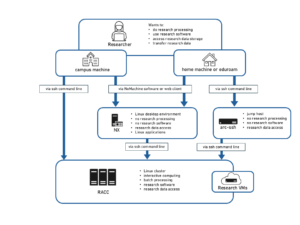
Matlab jobs can run in batch on the Reading Academic Computing Cluster (RACC) and the met-cluster, without the need for an interactive session or the Matlab user interface. We suggest you use the RACC as met-cluster is being decommissioned! In order to do that, an additional job script needs to be written, which calls the execution of the Matlab script after loading the Matlab module and finding the directory of the Matlab script.
The examples below show a batch job scripts that call a Matlab script (called testscript here), for the RACC:
#!/bin/bash #SBATCH --ntasks=1 #SBATCH --cpus-per-task=1 #(some Matlab jobs might benefit from getting more CPUs) #SBATCH --output=logfile.txt #SBATCH --mem=6G #(let's specify memory for the whole thing rather than per CPU) #SBATCH --time=120:00 # 2 hours module load matlab unset DISPLAY cd /path/to/your/matlab/script matlab -nodisplay < testscript.m > outfile.txt
and for met-cluster (legacy)
#!/bin/bash module load matlab cd /path/to/your/matlab/script matlab -nodisplay < testscript.m > outfile.txt
The ‘matlab’ command needs to be run with the ‘-nodisplay‘ flag in order to suppress the user interface. You can redirect the output into a file (called outfile.txt here), so that the progress of the Matlab job can be tracked easily, and is is separate from other output. But, if you don’t it will work fine too, and you get Matlab output in the job’s output file (logfile.txt). In Slurm, we need to redirect the input file in, ‘-r’ option, used in the met-cluster version of this article, might not work. Please note that the Matlab script itself should have the command ‘exit’ at the end so that Matlab exits successfully after it runs the code.
One known issue with the above script is that if you write/edit it through Windows, invisible extra characters tend to be introduced which stop the script from running. It’s a good idea to write this in the Unix environment instead.
You can execute the above script locally first before submitting it (remember to set the permissions of the script to ‘rx’ first).
On the RACC you submit the script with the command:
sbatch batch_job.sh
You will have to experiment to find how much memory the job will require. Allocating too much will be wasteful (and bad for your queuing priority), while allocating too little will get the job killed when it runs out of memory. Unfortunately Matlab often allocates a lot of memory even if your actual data does not use that much, and in Slurm you need to reserve memory for the whole thing. (Maybe we should test how it works with Matlab compiler at some point) It is safe to request just 1 CPU for the whole job (–cpus-per-task=1); if Matlab wants to run multi-threaded it will see how many CPUs it got and it will not attempt that (on met-cluster it is more tricky).
On met-cluster, the batch script can be submitted with the ‘qsub‘ command. To submit the job to e.g. the limited queue, you can use
qsub -pe smp 4 -l limited -j y -o $PWD/logfile.txt batch_job.sh
The flag ‘-j y -o $PWD/logfile.txt‘ pipes the system output from the batch jobs into one file in your run directory, which is useful for error tracking if your job crashes or fails to run. We added ‘-pe smp 4’. It is not needed for all Matlab jobs, but we mention it here because many Matlab jobs currently running on met-cluster are multi-threaded, while users are not aware that some Matlab functions used are are trying to parallelize the computations. Typically those jobs tend to use around 4 CPU cores,
For more information on how to use the clusters, please see the met-cluster documentation and Reading Academic Computing Cluster User’s Guide.
Compiled Matlab
It is also possible to compile your matlab scripts using the Matlab Compiler for the appropriate version of Matlab. This can considerably reduce the amount of memory required to run the script.
You can compile you matlab code using mcc. To access the compiler you need to setup MATLAB. This can be done, as before, with the command
module load matlab
If the file highlight.m contains the following lines
function result = highlight(a); result = strcat('**', a, '**')
Then it can be compiled with the command
mcc -R '-nodisplay,-nojvm,-singleCompThread' -m highlight.m
The options are:
-R - Runtime Matlab Options -nodisplay - Do not try to connect to an X11 display -nojvm - Do not use Java Virtual Machine -singleCompThread - Only a single thread, this allows PBS to run multiple copies of the job on one node. -m - Compile the code into a standalone executable
More information about the options can be found on the mcc page.
You can run the script using the command
./highlight $(hostname)
In your batch submission script you will need to load the same matlab module as you used to compile the code so that the environment is setup correctly.
#!/bin/bash #SBATCH --ntasks=1 #SBATCH --cpus-per-task=1 #(some Matlab jobs might benefit from getting more CPUs) #SBATCH --output=logfile.txt #SBATCH --mem=1G #(let's specify memory for the whole thing rather than per CPU) #SBATCH --time=120:00 # 2 hours module load matlab unset DISPLAY cd /path/to/your/matlab/compiled/script ./highlight $(hostname)
 Running Matlab Scripts as Batch Jobs
Running Matlab Scripts as Batch Jobs